| 规格 | 价格 | 库存 | 数量 |
|---|---|---|---|
| 5mg |
|
||
| 10mg |
|
||
| 50mg |
|
||
| 100mg |
|
||
| Other Sizes |
|
| 体外研究 (In Vitro) |
当酵母细胞暴露于 SMER3 (0-60 μM) 时,Met4 泛素化会受到抑制。 SMER3可以减轻met4Δ细胞的生长抑制[1]。通过以剂量依赖性方式将 SMER3 添加到连接酶过程中,可以减少 SCFMet30 对 Met4 的泛素化。 SMER3 不会改变 Skp1 或 Met30 蛋白的量,但会强烈抑制 Met30 与 Skp1 的结合 [1]。
|
|---|---|
| 参考文献 | |
| 其他信息 |
LSM-42773 是一种芳香酮。
|
| 分子式 |
C11H4N4O2
|
|---|---|
| 分子量 |
224.17
|
| 精确质量 |
224.033
|
| CAS号 |
67200-34-4
|
| PubChem CID |
568763
|
| 外观&性状 |
Light yellow to green yellow solid powder
|
| 密度 |
1.7±0.1 g/cm3
|
| 沸点 |
437.2±55.0 °C at 760 mmHg
|
| 熔点 |
296 °C(dec.)
|
| 闪点 |
218.2±31.5 °C
|
| 蒸汽压 |
0.0±1.0 mmHg at 25°C
|
| 折射率 |
1.779
|
| LogP |
0.9
|
| tPSA |
81.77
|
| 氢键供体(HBD)数目 |
0
|
| 氢键受体(HBA)数目 |
6
|
| 可旋转键数目(RBC) |
0
|
| 重原子数目 |
17
|
| 分子复杂度/Complexity |
350
|
| 定义原子立体中心数目 |
0
|
| InChi Key |
SFSSAKVWCKFRHE-UHFFFAOYSA-N
|
| InChi Code |
InChI=1S/C11H4N4O2/c16-9-6-4-2-1-3-5(6)7-8(9)13-11-10(12-7)14-17-15-11/h1-4H
|
| 化学名 |
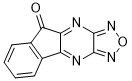
13-oxa-10,12,14,16-tetrazatetracyclo[7.7.0.02,7.011,15]hexadeca-1(16),2,4,6,9,11,14-heptaen-8-one
|
| 别名 |
SMER3 SMER 3 SMER-3
|
| HS Tariff Code |
2934.99.9001
|
| 存储方式 |
Powder -20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| 运输条件 |
Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs)
|
| 溶解度 (体外实验) |
DMSO : ~12.5 mg/mL (~55.76 mM)
|
|---|---|
| 溶解度 (体内实验) |
配方 1 中的溶解度: 1.25 mg/mL (5.58 mM) in 10% DMSO + 40% PEG300 +5% Tween-80 + 45% Saline (这些助溶剂从左到右依次添加,逐一添加), 悬浮液;超声助溶。
例如,若需制备1 mL的工作液,可将100 μL 12.5 mg/mL澄清的DMSO储备液加入到400 μL PEG300中,混匀;再向上述溶液中加入50 μL Tween-80 +,混匀;然后加入450 μL生理盐水定容至1 mL。 *生理盐水的制备:将 0.9 g 氯化钠溶解在 100 mL ddH₂O中,得到澄清溶液。 请根据您的实验动物和给药方式选择适当的溶解配方/方案: 1、请先配制澄清的储备液(如:用DMSO配置50 或 100 mg/mL母液(储备液)); 2、取适量母液,按从左到右的顺序依次添加助溶剂,澄清后再加入下一助溶剂。以 下列配方为例说明 (注意此配方只用于说明,并不一定代表此产品 的实际溶解配方): 10% DMSO → 40% PEG300 → 5% Tween-80 → 45% ddH2O (或 saline); 假设最终工作液的体积为 1 mL, 浓度为5 mg/mL: 取 100 μL 50 mg/mL 的澄清 DMSO 储备液加到 400 μL PEG300 中,混合均匀/澄清;向上述体系中加入50 μL Tween-80,混合均匀/澄清;然后继续加入450 μL ddH2O (或 saline)定容至 1 mL; 3、溶剂前显示的百分比是指该溶剂在最终溶液/工作液中的体积所占比例; 4、 如产品在配制过程中出现沉淀/析出,可通过加热(≤50℃)或超声的方式助溶; 5、为保证最佳实验结果,工作液请现配现用! 6、如不确定怎么将母液配置成体内动物实验的工作液,请查看说明书或联系我们; 7、 以上所有助溶剂都可在 Invivochem.cn网站购买。 |
| 制备储备液 | 1 mg | 5 mg | 10 mg | |
| 1 mM | 4.4609 mL | 22.3045 mL | 44.6090 mL | |
| 5 mM | 0.8922 mL | 4.4609 mL | 8.9218 mL | |
| 10 mM | 0.4461 mL | 2.2305 mL | 4.4609 mL |
1、根据实验需要选择合适的溶剂配制储备液 (母液):对于大多数产品,InvivoChem推荐用DMSO配置母液 (比如:5、10、20mM或者10、20、50 mg/mL浓度),个别水溶性高的产品可直接溶于水。产品在DMSO 、水或其他溶剂中的具体溶解度详见上”溶解度 (体外)”部分;
2、如果您找不到您想要的溶解度信息,或者很难将产品溶解在溶液中,请联系我们;
3、建议使用下列计算器进行相关计算(摩尔浓度计算器、稀释计算器、分子量计算器、重组计算器等);
4、母液配好之后,将其分装到常规用量,并储存在-20°C或-80°C,尽量减少反复冻融循环。
计算结果:
工作液浓度: mg/mL;
DMSO母液配制方法: mg 药物溶于 μL DMSO溶液(母液浓度 mg/mL)。如该浓度超过该批次药物DMSO溶解度,请首先与我们联系。
体内配方配制方法:取 μL DMSO母液,加入 μL PEG300,混匀澄清后加入μL Tween 80,混匀澄清后加入 μL ddH2O,混匀澄清。
(1) 请确保溶液澄清之后,再加入下一种溶剂 (助溶剂) 。可利用涡旋、超声或水浴加热等方法助溶;
(2) 一定要按顺序加入溶剂 (助溶剂) 。
|
|
|